The Tulane Center for Translational Research in Infection & Inflammation | NextGen Sequencing Core at Tulane University is excited to start using, iLAB, an online system to streamline the process of requesting and billing for core services. All facility users are invited to use the system, which requires a one-time registration. Once registered, the system will enable you to place service requests, provide required approvals, and monitor progress.
To Register for an iLab Account:
- Complete the registration form on the sign-up page.
- Receive a Welcome Email from iLab (typically within one business day) with login credentials.
To Create a Service Request:
Once you have been accepted into your PI’s lab and assigned a Fund Number, you can submit requests.
- Navigate to the core page:
https://my.ilabsolutions.com/service_center/show_external/6248
- In the upper-right-hand corner click ‘Sign In’ A pop-up window should appear displaying, ‘Sign in using iLab credentials.’ Click the iLab link.
- Enter the credentials received in your welcome email from iLab.
- Select the Request Services tab and click on the ‘Request Service’ button next to the service of interest.
- You will be asked to complete a form and provide payment information for your request before submitting the request to the core.
- Your request will be pending review by the core. The core will add charges and submit it back to you for approval. Make sure to watch for an email from iLab regarding your updated Fund.
Additional Help
More detailed instructions are available on our helpsite. For any questions not addressed, click on the “HELP” link in the upper right hand corner or contact ilab-support@agilent.com.
Purpose
The Center for Transcriptional Research in Infection and Inflammation offers Illumina Next Generation sequencing services to support NGS-based research studies of gene expression as well as single-cell RNAseq to the Tulane University research community.
The Sequencing Core has an interactive staff, dedicated to the production and delivery of high-quality sequence data.
QC Instrumentation
Quantification: Invitrogen Qubit
- More sensitive than UV absorbance
- Uses 1 uL of sample
- Fast, reliable
Qualification: Agilent TapeStation 4150
- Automated electrophoresis for Sample Size and Integrity
- Constant cost per sample
- Uses 2ul of sample
- Fast, reliable
10X Chromium Single Cell
Complete workflow from cell suspension through sequencing-ready cDNA library.

- Flexible throughput encapsulates 100 – 80,000+ cells in 10 minutes
- Cell capture efficiency up to 65%
- Compatible with Illumina® sequencers
- More efficient than leading academic droplet systems
Illumina NextSeq 2000

- The NextSeq 2000 System brings the power of a high throughput sequencing system to your bench top. With tunable output and high data quality, it provides the flexible power you need for whole genome, transcriptome, and targeted resequencing.
- NextSeq 2000 Systems support emerging and mid throughput sequencing applications as well as a broad range of methods such as exome sequencing, target enrichment, single cell profiling, transcriptome sequencing, and more. They offer an intuitive workflow with load and go ease and visual cues about run status.
Bioinformatic Resources
Cypress is Tulane's newest HPC cluster, offered by Technology Services for use by the Tulane research Community. It is a 124-node cluster, with each node providing dual 10-core 2.8 GHz Intel Xeon E5-2680 v2 CPUs, 64 GB of RAM, and dual Xeon Phi 7120P coprocessors. Nodes are interconnected on a 40 Gigabit Ethernet network using a single Dell Z9500 Ethernet Fabric Switch.
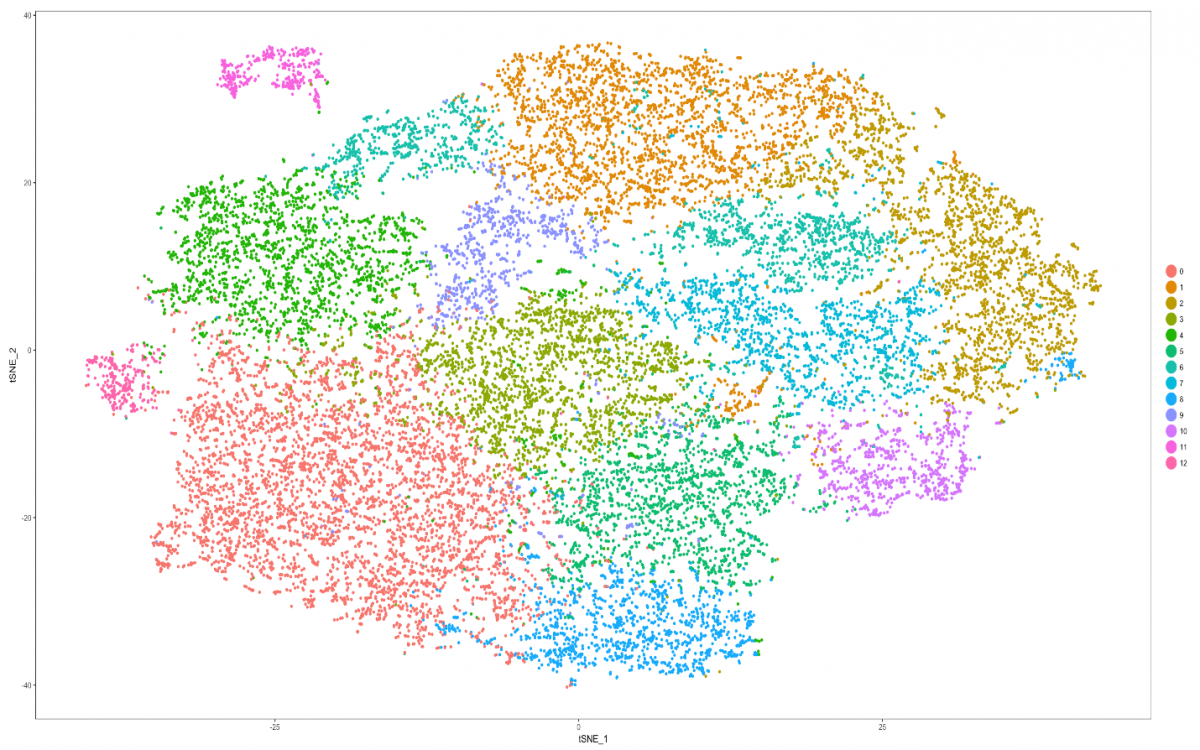
Cell Ranger V3.0 by 10x Genomics is run on cypress. Cell Ranger is a set of analysis pipelines that process Chromium single-cell RNA-seq output to align reads, generate feature-barcode matrices and perform clustering and gene expression analysis.” Cell Ranger data can be viewed interactively with the 10x Loupe Browser. Subsequent analysis is done in R using Seurat and Monocle packages.
Seurat is an R package designed for QC, analysis, and exploration of single-cell RNA-seq data. Seurat aims to enable users to identify and interpret sources of heterogeneity from single-cell transcriptomic measurements, and to integrate diverse types of single-cell data.
Monocle provides tools for analyzing single-cell expression experiments. Monocle introduced the strategy of ordering single cells in pseudotime, placing them along a trajectory corresponding to a biological process such as cell differentiation by taking advantage of individual cell's asynchronous progression of those processes.
The Sequencing Core is located on the 3rd floor of J. Bennett Johnston Building, Room 339 in the Center for Translational Research in Infection and Inflammation.
The core is currently accepting samples, for more information reach out to Jay Kolls or Kejing Song.
CONTACT
Jay K. Kolls, MD
Center for Translational Research
in Infection and Inflammation
Phone: 504-988-0456
Email: jkolls1@tulane.edu
For more information:
Administrative/Financial: Candice Fisher cfisher9@tulane.edu
Single Cell (10X): Kejing Song ksong1@tulane.edu
Bulk RNA Seq: Haiyan Miller hdeng@tulane.edu
Bioinformatics: MST Shamima Khatun, Ph.D. mkhatun@tulane.edu
